Data classes#
Experiment#
An Experiment is set of movies which are meant to be analyzed together.
This class scans all .tif files in data_folder and stores localizations and tracks generated by pipelines in the export_folder.
exp = Experiment(data_folder="myproject/data/raw",export_folder="myproject/data/processed")
Important
Pixel size and camera exposure are assumed constant across all movies of an experiment.
The first time you create an Experiment with a given export_folder, you will be prompted to enter the corresponding values.
They will then be stored in the export folder and you will not have to enter them again.
Process batches of movies#
Once you have set up your ImagePipeline, you might want to run it on all your movies. To do so, simply run the following command.
This will create in your experiment’s directory a folder named after yout pipeline’s name.
The processing outputs will be written in this folder and loaded when needed.
The pipeline’s parameters will be saved in your Experiment’s export_folder.
tp.process(exp)
The index table#
An experiment stores movie-specific information in an index table (index_df) in order to be accessible for later analysis :
which type of cell did you observe, which protein did you look at, how were the cells treated, etc…
Below is an example of what such a file might look like.
file |
cell_type |
observation_time |
|---|---|---|
ROI1.tif |
neuron |
T+1h |
ROI2.tif |
astrocyte |
T+1h |
ROI3.tif |
neuron |
T+2h |
Populate columns from file names#
This information can be entered manually, by editing the corresponding .csv file
or by overriding the custom_fields property of a subclass of Experiment to
automatically populate the columns from the file names. It is a dictionnary whose keys are the desired column names and whose values indicate how to populate the rows.
If the value is an integer
i, the column’s rows will take the value of thei-th part of the file path (note that it can be negative, if you’d like to start from the end).If it is a string, the rows will take the value
Trueif the file path contains this string, andFalseotherwise.Finally, if it is a callable taking one argument, then rows’ values will be
callable(file_path).
from palmari import Experiment
def myfunc(s:str):
return s.split("_")[4].upper()
class MyExperiment(Experiment):
@property
def custom_fields(self) -> dict:
return {
"field_1":"sometext", # True if the file's name contains "sometext"
"field_2":3, # returns the third part of the file name (delimited by the filesystem's separator)
"field_3":myfunc # Returns the output of myfunc
}
Tip
Access the file names of an experiment’s movies by treating the experiment as a list.
for file_path in exp:
# do stuff
# or
my_file = exp[0]
Acquisition#
Each movie in an Experiment is an instance of the Acquisition class.
It must be instanciated with a reference to a ImagePipeline responsible for generating or loading its localizations.
acq = Acquisition(image_file=exp[0], experiment=exp, image_pipeline=tp)
# To process a single acquisition with ImagePipeline tp, use the following function
tp.process(acq)
If the movie has been processed by the given pipeline, then you can access its localizations table using .locs.
Here is what this table looks like after an acquisition has been processed by a standard pipeline.
The n column identifies the trajectory to which the localization belongs.
Time is expressed in seconds and coordinates are in micrometers.
frame |
x |
y |
ratio |
sigma |
total_intensity |
t |
n |
|
|---|---|---|---|---|---|---|---|---|
0 |
0 |
4.634299 |
2.957245 |
1.140336 |
4.373642 |
3572.0 |
0.00 |
0 |
1 |
0 |
5.133801 |
5.043820 |
1.003860 |
4.088229 |
3611.0 |
0.00 |
1 |
2 |
2 |
4.424841 |
5.378470 |
1.337645 |
4.451456 |
3790.0 |
0.06 |
2 |
3 |
2 |
5.673679 |
2.878053 |
1.114035 |
4.624364 |
4374.0 |
0.06 |
3 |
4 |
3 |
4.039363 |
4.773757 |
1.458809 |
3.972434 |
5811.0 |
0.09 |
2 |
After the movie has been localized and tracked,
you can visualize the localizations superimposed to the camera recording in Napari using the .view() function.
acq.view() # View the .tif movie with superimposed localizations and tracks, using Napari.
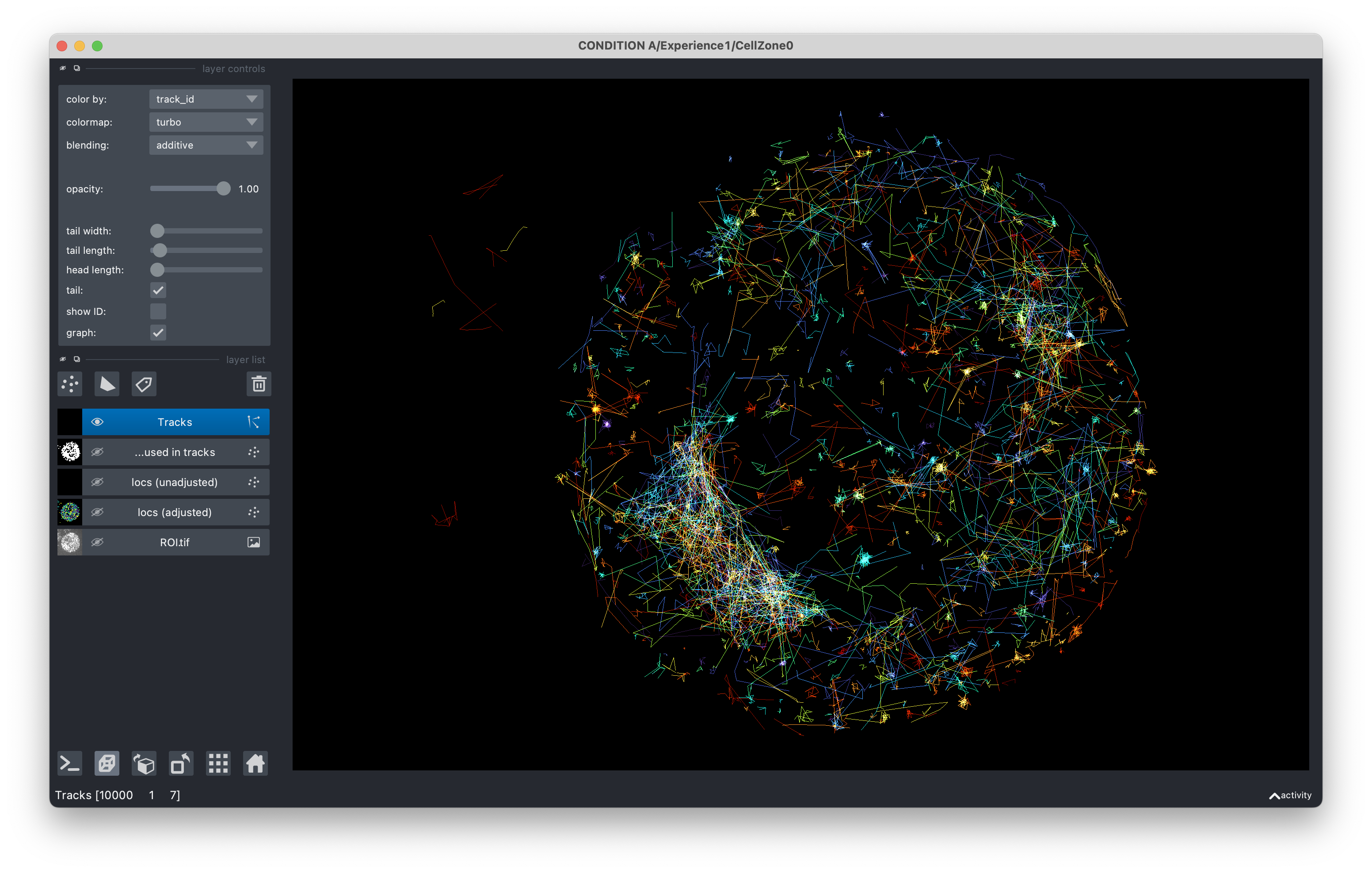
There you are !#